-Search query
-Search result
Showing 1 - 50 of 1,137 items for (author: joseph & n)

EMDB-42070: 
DpHF18 filament
Method: helical / : Lynch EM, Shen H, Kollman JM, Baker D

EMDB-42075: 
DpHF7 filament
Method: helical / : Lynch EM, Farrell D, Shen H, Kollman JM, DiMaio F, Baker D

EMDB-42088: 
DpHF19 filament
Method: helical / : Lynch EM, Shen H, Kollman JM, Baker D

PDB-8uao: 
DpHF18 filament
Method: helical / : Lynch EM, Shen H, Kollman JM, Baker D

PDB-8ub3: 
DpHF7 filament
Method: helical / : Lynch EM, Farrell D, Shen H, Kollman JM, DiMaio F, Baker D

PDB-8ubg: 
DpHF19 filament
Method: helical / : Lynch EM, Shen H, Kollman JM, Baker D

EMDB-17329: 
48S late-stage initiation complex with m6A mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

EMDB-17330: 
48S late-stage initiation complex with non methylated mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

PDB-8p03: 
48S late-stage initiation complex with m6A mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

PDB-8p09: 
48S late-stage initiation complex with non methylated mRNA
Method: single particle / : Guca E, Lima LHF, Boissier F, Hashem Y

EMDB-41816: 
Cryo-EM structure of the RAF1-HSP90-CDC37 complex in the closed state
Method: single particle / : Finci LI, Simanshu DK

EMDB-41817: 
Cryo-EM structure of the HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

EMDB-41818: 
Cryo-EM structure of the cross-linked HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

PDB-8u1l: 
Cryo-EM structure of the RAF1-HSP90-CDC37 complex in the closed state
Method: single particle / : Finci LI, Simanshu DK

PDB-8u1m: 
Cryo-EM structure of the HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

PDB-8u1n: 
Cryo-EM structure of the cross-linked HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK
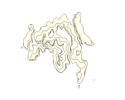
EMDB-19184: 
Late alpha-Synuclein fibril structure from liquid-liquid phase separations.
Method: helical / : De Simone A, Barritt JD, Chen S, Cascella R, Cecchi C, Bigi A, Jarvis JA, Chiti F, Dobson CM, Fusco G

PDB-8ri9: 
Late alpha-Synuclein fibril structure from liquid-liquid phase separations.
Method: helical / : De Simone A, Barritt JD, Chen S, Cascella R, Cecchi C, Bigi A, Jarvis JA, Chiti F, Dobson CM, Fusco G
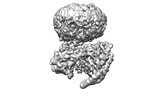
EMDB-42023: 
GPR3 Orphan G-coupled Protein Receptor in complex with Dominant Negative Gs.
Method: single particle / : Russell IC, Belousoff MJ, Sexton P

PDB-8u8f: 
GPR3 Orphan G-coupled Protein Receptor in complex with Dominant Negative Gs.
Method: single particle / : Russell IC, Belousoff MJ, Sexton P

EMDB-19038: 
Structure of Integrator-PP2A bound to a paused RNA polymerase II-DSIF-NELF-nucleosome complex
Method: single particle / : Fianu I, Ochmann M, Walshe JL, Cramer P

EMDB-19040: 
Structure of Integrator-PP2A-SOSS-CTD post-termination complex
Method: single particle / : Fianu I, Ochmann M, Walshe JL, Cramer P

EMDB-19047: 
Structure of Integrator-PP2A complex
Method: single particle / : Fianu I, Ochmann M, Walshe JL, Cramer P

PDB-8rbx: 
Structure of Integrator-PP2A bound to a paused RNA polymerase II-DSIF-NELF-nucleosome complex
Method: single particle / : Fianu I, Ochmann M, Walshe JL, Cramer P

PDB-8rbz: 
Structure of Integrator-PP2A-SOSS-CTD post-termination complex
Method: single particle / : Fianu I, Ochmann M, Walshe JL, Cramer P

PDB-8rc4: 
Structure of Integrator-PP2A complex
Method: single particle / : Fianu I, Ochmann M, Walshe JL, Cramer P

EMDB-41805: 
Cryo-EM structure of murine Thrombopoietin receptor ectodomain in complex with Tpo
Method: single particle / : Sarson-Lawrence KS, Hardy JM, Leis A, Babon JJ, Kershaw NJ

PDB-8u18: 
Cryo-EM structure of murine Thrombopoietin receptor ectodomain in complex with Tpo
Method: single particle / : Sarson-Lawrence KS, Hardy JM, Leis A, Babon JJ, Kershaw NJ

EMDB-43542: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation
Method: single particle / : Petroff II JT, Deng Z, Rau MJ, Fitzpatrick JAJ, Yuan P, Cheng WWL

PDB-8vuw: 
ELIC5 with cysteamine in 2:1:1 POPC:POPE:POPG nanodisc in open conformation
Method: single particle / : Petroff II JT, Deng Z, Rau MJ, Fitzpatrick JAJ, Yuan P, Cheng WWL

EMDB-18245: 
Plunge-frozen (control) map of beta-galactosidase
Method: single particle / : Esser TK, Boehning J, Bharat TAM, Rauschenbach S

EMDB-41986: 
Human retinal variant phosphomimetic IMPDH1(595)-S477D free octamer bound by GTP, ATP, IMP, and NAD+
Method: single particle / : Calise SJ, Kollman JM

EMDB-41989: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by GTP, ATP, IMP, and NAD+, octamer-centered
Method: single particle / : Calise SJ, Kollman JM

EMDB-42012: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by GTP, ATP, IMP, and NAD+, interface-centered
Method: single particle / : Calise SJ, Kollman JM

EMDB-42026: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by ATP, IMP, and NAD+, octamer-centered
Method: single particle / : Calise SJ, Kollman JM

EMDB-42029: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by ATP, IMP, and NAD+, interface-centered
Method: single particle / : Calise SJ, Kollman JM

PDB-8u7m: 
Human retinal variant phosphomimetic IMPDH1(595)-S477D free octamer bound by GTP, ATP, IMP, and NAD+
Method: single particle / : Calise SJ, Kollman JM

PDB-8u7q: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by GTP, ATP, IMP, and NAD+, octamer-centered
Method: single particle / : Calise SJ, Kollman JM

PDB-8u7v: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by GTP, ATP, IMP, and NAD+, interface-centered
Method: single particle / : Calise SJ, Kollman JM

PDB-8u8o: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by ATP, IMP, and NAD+, octamer-centered
Method: single particle / : Calise SJ, Kollman JM

PDB-8u8y: 
Human retinal variant phosphomimetic IMPDH1(546)-S477D filament bound by ATP, IMP, and NAD+, interface-centered
Method: single particle / : Calise SJ, Kollman JM

EMDB-36766: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36767: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36768: 
20 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36769: 
30 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36770: 
40 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36771: 
50 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36772: 
60 Degree Tilted Single-Particle CryoEM Reconstruction of AAV2 Viral Capsid
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36807: 
Untilted (0 Degree Tilted) Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D

EMDB-36809: 
10 Degree Tilted Single-Particle CryoEM Reconstruction of Apoferritin
Method: single particle / : Aiyer S, Tan YZ, Lyumkis D
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
